
Cilia are tiny and hair-like organelles extruding from the surface of most cell types, serving either sensory or motile functions. In the eyes, the photoreceptor cells possess sensory cilia that are important for vision. Motile cilia align along the surface of the respiratory tract and function in airway clearance of mucus and inhaled pathogens. Mutations in retinitis pigmentosa GTPase regulator (RPGR) affect both photoreceptor sensory cilia and airway motile cilia, leading to retinitis pigmentosa and the rare motile ciliopathy primary ciliary dyskinesia (PCD), respectively. Not all patients develop PCD, and currently, it remains unclear which RPGR variants predispose patients to PCD.
To elucidate how loss of RPGR impacts motile cilia and contributes to respiratory disease, the research team led by Prof. Zhen LIU from the Division of Life Science at the Hong Kong University of Science and Technology (HKUST) leverages organoids, super-resolution microscopy, and live-cell imaging to investigate the nasal multiciliated cells derived from patients carrying RPGR variants, as well as CRISPR-engineered RPGR knockout multiciliated cells. The study, recently published in the Journal of Clinical Investigation, provides new mechanistic insights into this disease.
In collaboration with physicians from the Hospital for Sick Children and BC Children’s Hospital, Canada, the team analyzed a cohort of 32 patients with different pathological RPGR variants and found that defective and disorganized ciliary structures resulted in impaired ciliary beating or beat coordination. To inspect how RPGR regulates motile cilia, they utilized different super-resolution microscopy modalities and revealed the abnormal condensed apical F-actin meshwork in both patient-derived and RPGR KO multiciliated cells. Critically, these ciliary abnormalities can be ameliorated by treatments that disrupt the accumulated F-actin.
This study uncovers a distinct role of RPGR in regulating F-actin dynamics at the apical surface, thereby coordinating multiciliogenesis and maintaining proper ciliary beating. The methodologies and findings have potential clinical applications in diagnosing this rare disease and improving patient outcomes with therapeutic interventions. This work closely supports HKUST’s newly established School of Medicine and reflects the University’s increasing focus on translational medicine.
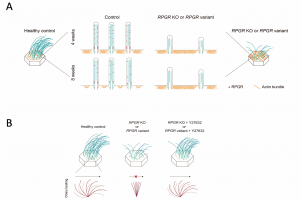
The multiciliated cells from patients with RPGR variants have spare, short, and mostly static cilia. These defects can be ameliorated by treatments that disrupt F-actin accumulation.
The link to the paper: https://www.jci.org/articles/view/193367
Read More
The Innovation and Technology Commission of the HKSAR Government has recently announced the results of the third round of projects selected for funding under the Research, Academic and Industry Sectors One‑plus (RAISe+) Scheme.
Congratulations to Project Leader Prof. ZHANG Lisheng, (Assistant Professor in the Division of Life Science and the Department of Chemistry), whose project has been selected for funding: “RNA Epitranscriptomics and AI: Empowering First‑Strike Screening to Prevent Cancer Before Its Onset”.
Prof. ZHANG’s research focuses on a “first-strike” approach to oncology. By combining the study of RNA epitranscriptomics (the biochemical modifications of RNA) with Artificial Intelligence (AI), the project aims to detect the very first molecular signatures of cancer. This technology could allow for screening and intervention long before physical symptoms or tumors appear, potentially saving countless lives through ultra-early prevention.

Research teams from the Division of Life Science (LIFS) at The University of Science and Technology (HKUST) achieved recognition at the 51st International Exhibition of Inventions Geneva (Geneva Inventions Expo).
HKUST’s 62 participating teams won a total of 62 accolades–including 13 Gold Medals with Congratulations of the Jury, 20 Gold Medals, 20 Silver Medals, and 9 Bronze Medals. This record-breaking performance distinguishes HKUST as the highest-awarded higher education institution in Hong Kong at this year’s Expo.
Principal Investigators: Prof. ZHU Guang, (Professor, Division of Life Science, and Department of Chemical and Biological Engineering); Dr. XU Naining, (Post-doctoral Fellow, LIFS)
The Challenge: According to the World Health Organization (WHO), an estimated 5.4 million people are bitten by snakes annually, resulting in many cases of envenoming. This leads to approximately 81,000 to 138,000 deaths each year. In Hong Kong, the Poison Control Centre reported nearly 300 snakebite‑related hospital admissions between 2021 and 2023. Traditional antivenom production involves injecting snake venom into large animals, such as horses or sheep, to stimulate antibody production. The antibodies are then extracted, a costly process that may cause allergic reactions in some individuals.
The Innovation: The team’s next‑generation recombinant antivenom utilizes human monoclonal antibody cocktails developed through a single B-cell screening platform to significantly reduce allergic responses. Lightweight and easy to use, this autoinjector enables immediate on‑site use with a single press to save critical time during emergencies.
Principal Investigator: Prof. ZHANG Li-Sheng, (Assistant Professor, Division of Life Science and Department of Chemistry) with CHAN Hei-Man and TSE Man–Hin (MPhil students, LIFS).
The Challenge: Colorectal cancer is the second leading cause of cancer deaths in Hong Kong, with a five‑year survival rate of only about 9.3% for late‑stage cases. However, colonoscopy, the clinical gold standard, is invasive, leading to low participation rates. In addition, stool‑based tests and blood‑derived cell-free DNA assays often lack the necessary sensitivity to detect precancerous lesions and Stage 0 cancers, which can delay diagnoses that could have been prevented.
The Innovation: The team has developed a novel blood screening platform, EpiLumenix, which integrates cell‑free RNA epitranscriptomics and AI to capture dynamic tumor microenvironment fingerprints from just 2 mL of blood. This platform achieves a groundbreaking 75% sensitivity for Stage 0 colorectal cancer and advanced adenomas, significantly outperforming current commercial stool and cfDNA tests. As a less‑invasive solution, it requires no fasting or bowel preparation, thereby improving patient compliance, and shifting colorectal cancer management from passive treatment to proactive, precision-based early detection and prevention.
Targeting C9orf72 G-Quadruplexes: Structure-Based and AI-Optimized Small Molecule Therapeutics for ALS/FTD
Principal Investigators: Prof. ZHU Guang, (Professor, LIFS, and CBE); Dr. XU Naining, (Post-doctoral Fellow, LIFS)
The Challenge: Amyotrophic Lateral Sclerosis (ALS) is a fatal neurodegenerative disorder characterized by the progressive loss of motor neurons. Patients face a devastating prognosis, with a median survival of only 2 to 5 years following diagnosis, and no curative therapies currently available. A major genetic contributor to ALS is the C9orf72 mutation, which involves extensive G4C2 DNA/RNA repeat expansions that form toxic aggregates and drive neuronal dysfunction. As the global incidence of ALS increases with an aging population, existing FDA approved treatments remain largely symptomatic and fail to address this underlying genetic pathology. Consequently, there is a critical unmet medical need for a true disease modifying therapy that targets the root cause of ALS.
The Innovation: Led by Prof. ZHU Guang at HKUST, the research team developed GY 368, a first in class small molecule therapy that directly targets the C9orf72 gene mutation. Using an AI driven targeted drug discovery platform, the team optimized GY 368 to selectively resolve pathogenic G quadruplex structures. In preclinical models, GY 368 demonstrated robust blood–brain barrier penetration, significantly reduced toxic RNA foci, and improved motor function without observable toxicity. By addressing a fundamental genetic cause of ALS and frontotemporal dementia (FTD), this breakthrough therapy offers new promise for extending patient survival.
These achievements underscore HKUST’s commitment to translational research, bridging the gap between laboratory discovery and real-world industrial application to improve global healthcare outcomes and drive innovation in the biotechnology sector.
For further information please visit: HKUST News-Geneva Inventions Expo
Read More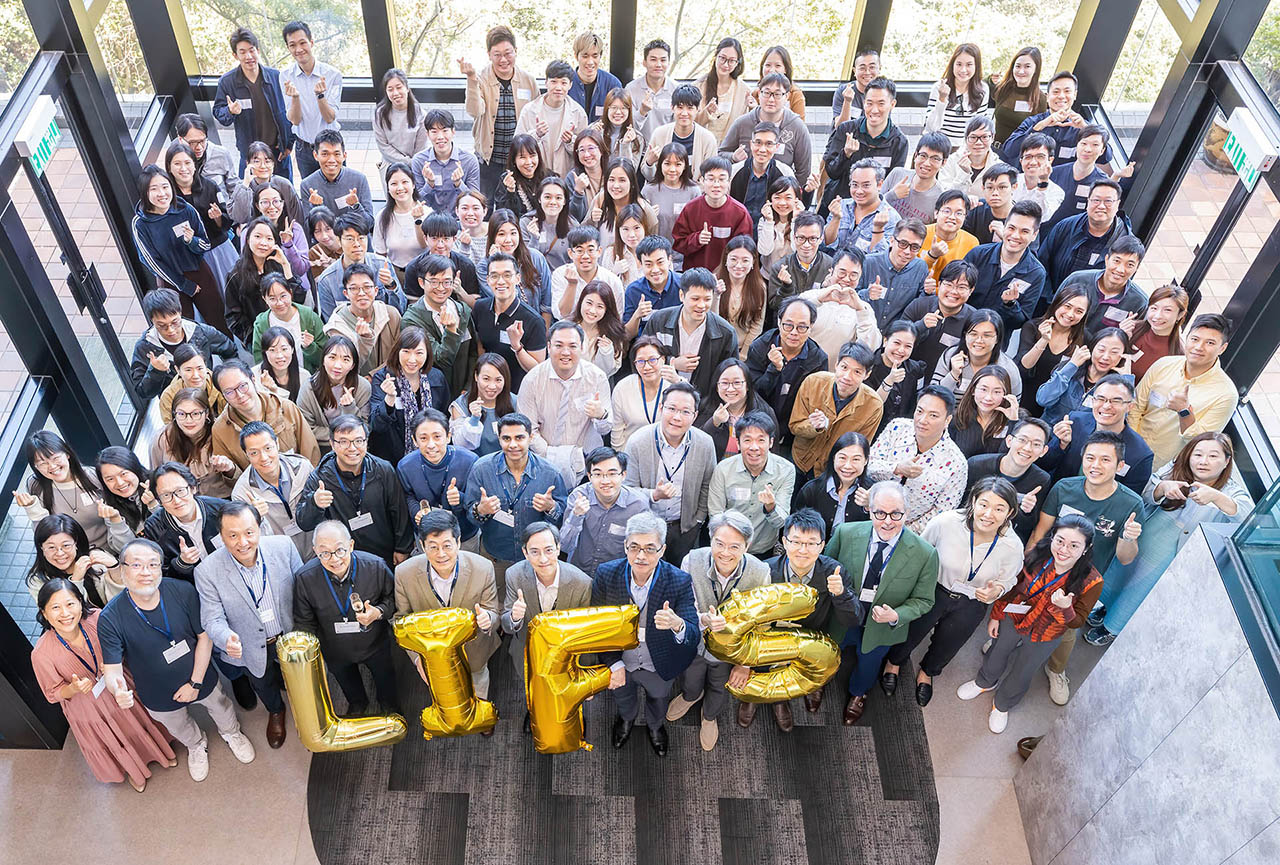
More than 90 alumni from the Division returned to campus on 14 March 2026 to reconnect with fellow graduates and professors at the LIFS Alumni Homecoming 2026.
The event started with a welcoming cocktail reception followed by a buffet lunch. Prof. Guojun BU, Head of Division, shared recent developments and achievements of LIFS, while Prof. Yung Hou WONG, Dean of Science, provided updates on the progress and future plans of HKUST’s Medical School.
Representatives from the Life Science Alumni Association (LSAA) introduced their activities and upcoming plans. Two new initiatives from the Development & Alumni Office (DAO) – Alumni Commons and the Class Ambassador Program – were also presented. Faculty members then led a celebratory toast before group photos were taken.

In the afternoon, alumni visited the newly launched Alumni Commons, designed to enhance alumni engagement and community building.
The gathering served as a meaningful opportunity for alumni and faculty to reconnect, build new ties, and stay updated with the Division’s latest developments. LIFS looks forward to continuing such reunions to strengthen alumni bonds and foster a lasting sense of belonging.
Repost from: https://alum.hkust.edu.hk/events/lifs-alumni-homecoming-2026
Read More
A research team from The Hong Kong University of Science and Technology (HKUST) has made a breakthrough discovery in understanding the molecular machinery of RNA silencing. The team uncovered how the human enzyme DICER achieves highly precise processing of microRNAs (miRNAs), advancing gene regulation research and offering new insights into the mechanisms underlying cancer, immune disorders, and genetic diseases.
The message of human life is encoded in our genomic DNA through transcription of messenger RNAs which carries and executes the genetic instructions. RNA molecules, typically single-stranded and composed of ribonucleotides (A, U, G, and C), play essential cellular roles ranging from protein synthesis and gene regulation to serving as genetic material in certain viruses. Within this RNA-based regulatory landscape, the enzyme DICER functions as a highly precise “molecular scissors.” It cleaves double-stranded RNA precursors into short regulatory RNAs that are subsequently incorporated into the RNA-induced silencing complex (RISC), enabling the cell to identify and suppress incorrect or unnecessary genetic messages—much like marking and deleting errors in a text.
For years, researchers have sought to understand how DICER achieves its extraordinary cleavage accuracy. Using state-of-the-art biochemical and structural biology techniques and high-resolution cryoelectron microscopy (cryo-EM), the HKUST research team visualized DICER–RNA interactions at an atomic level. The study’s first author Minh Khoa Ngo explained, “CryoEM allowed us to observe how RNA substrates engage with DICER at an atomic detail. These structural snapshots vividly reveal the dynamic adjustments DICER makes when processing different RNA sequences, fundamentally reshaping our understanding of how this enzyme functions.”
The team discovered that before cleavage occurs, DICER undergoes conformational adjustments that guide RNA substrates into the correct register. The enzyme then uses specific structural elements—particularly amino acids within 5′-end binding pockets—to align the RNA precisely before adopting the “cleavage ready” conformation.
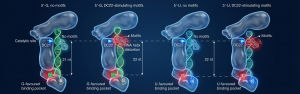
This diagram shows DICER’s Decision Process, illustrating how the enzyme acts like a “smart ruler” to decide exactly where to snip the RNA strand. The “G” Path (Left): When RNA starts with the letter G, it fits into a green pocket. Normally, this leads to a shorter cut (21 units). However, if the RNA has a specific internal feature, the enzyme physically twists the strand (red lines), overriding the rule to make a longer cut (22 units). The “U” Path (Right): When RNA starts with the letter U, it fits into a pink pocket. This pocket is shaped to naturally align the RNA for the longer cut (22 units) every time, regardless of its internal feature.
Prof. Nguyen, the corresponding author, elaborated: “It is as if the scissors can ‘read’ exactly where the RNA should be cut at single nucleotide resolution, ensuring the integrity of the entire message. Our study uncovers not only the previously known U-favoured 5′-end binding pocket, but also a newly identified G-favoured 5′-end binding pocket. Together, these form a dual-pocket mechanism that determines cleavage positioning, providing an entirely new framework for understanding how DICER accommodates and processes diverse RNA substrates.”
He further added: “The importance of this discovery extends beyond basic biology. By revealing how DICER integrates 5′-end identity, RNA motifs, and domain motions to maintain cleavage fidelity, our findings lay a mechanistic foundation for improving RNA-based therapeutics, optimizing gene silencing technologies, and uncovering the molecular origins of DICER-related genetic diseases.”
Repost from
https://hkust.edu.hk/news/hkust-researchers-achieve-breakthrough-rna-silencing-mechanism
Read More
Division of Life Science
LIFS Summer Recruitment Camp 2026
For 2027/28 Fall Postgraduate Admission
6 July 2026 – 10 July 2026 (5 days)
The Hong Kong University of Science and Technology (HKUST) promotes a vibrant and dynamic environment that emphasizes academic excellence, innovation and collaboration. Division of Life Science (LIFS) is host to cutting-edge, international quality research. There are six major areas of research interest including neuroscience, structural biology, cell & developmental biology, biotechnology, genomics and other interdisciplinary research.
Program & Highlights
This is a 5-day program for those who are interested in pursuing MPhil or PhD programs in Division of Life Science at HKUST in the Fall 2027/28.
Participants will attend a series of informal seminars introducing the current research topics and postgraduate programs offered in the Division of Life Science. These seminars are intended to foster discussion among the participants and faculty members and to serve as a basis for further social and scientific interactions. Participants will also have opportunities to visit the individual research labs and university research facilities, meeting with professors and fellow postgraduate students, and discuss research topics and others, such as preparing for graduate school.
The Program provides opportunity for the participants to learn the individual research groups and let the professors access your ability to conduct research projects in their MPhil or PhD programs.
Eligibility
Accommodation and Meals
Other fees
How to Apply?

The Summer Research Program is designed for outstanding end-of-year 3 undergraduate students who are interested in pursuing research postgraduate study in the Division of Life Science at the Hong Kong University of Science and Technology (HKUST). This program offers participants the unique opportunity to immerse themselves in the research life at HKUST campus and work with faculty mentors from our Division in conducting research projects in an interactive environment. Participants can also choose to join the interview session of the LIFS Summer Recruitment Camp, which will be held on July 8-9.
2 July – 6 August 2026
Undergraduate students studying in Mainland China and overseas, majoring in Life Science, Chemistry, Biotechnology, Bioengineering, Computational Biology and related programs, who have completed 3 years of study (out of a 4-year bachelor program) and have attained a GPA of 3.2 out of 4.0 (or 80%) or higher.
A subsidy of around HK$10,000 will be provided to cover the on-campus student hostel fee and other local expenses. To enroll in this Summer Research Program, participants are required to pay the summer internship application fee, insurance, visa fee and a 1-credit tuition fee in advance (approximately HK$3,300-HK4,000). Additionally, participants are responsible for applying for a visa (which may take 10-12 weeks) and covering their own transportation costs to and from Hong Kong.
Submit online application via the link:
https://ust.az1.qualtrics.com/jfe/form/SV_3Ld9WNQJimmOZLM
23 March 2026 (Monday)
The review of applications will be conducted by the Division of Life Science. Successful applicants will be notified by the Division as soon as the application is closed. After receiving the Division’s confirmation:
(For successful applicants only) Submit online Visiting Interns application to the Undergraduate Recruitment and Admissions Office (https://join.ust.hk/admissions/visiting/ ).
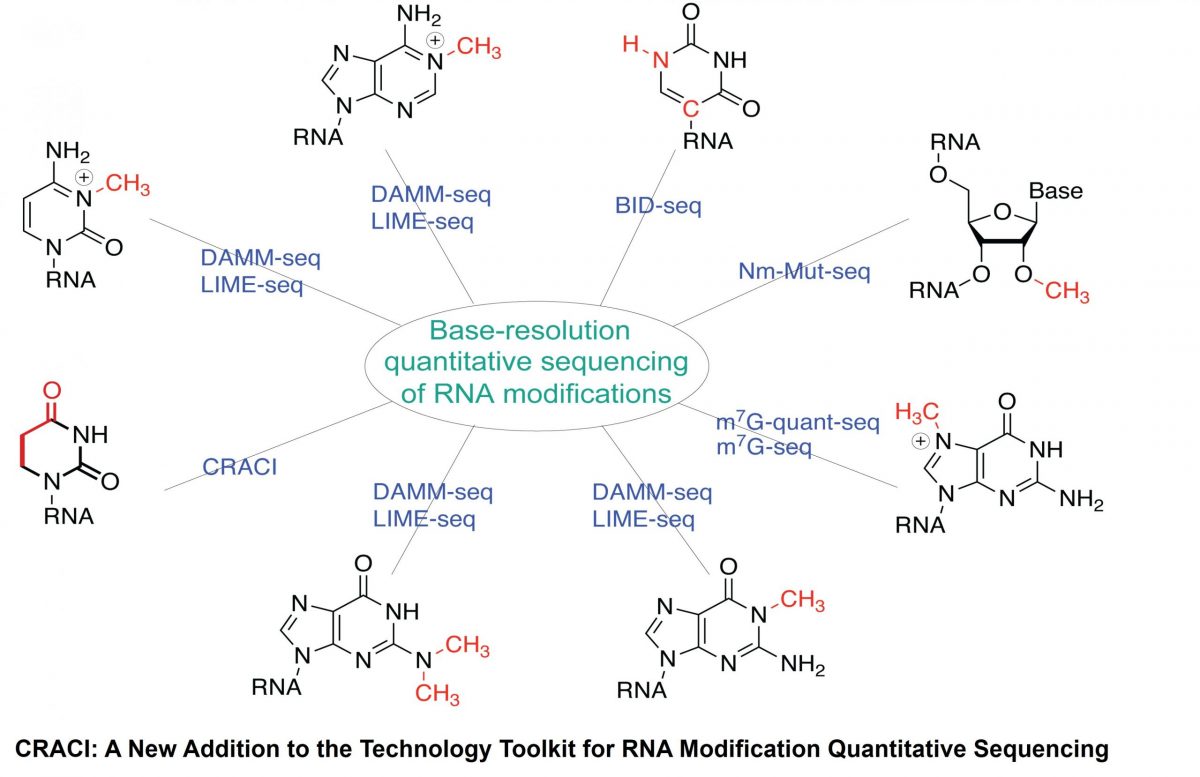
A team led by Prof. ZHANG Li‑Sheng (Division of Life Science & Department of Chemistry, HKUST) has developed Chemical Reduction Assisted Cytosine Incorporation sequencing (CRACI) — a sensitive method that quantitatively pinpoints dihydrouridine (D), the most abundant tRNA modification at the single-base resolution.
Small chemical marks on RNA—like D—can work like fine‑tuning structural switches, influencing how genes are expressed. Until now, these D marks were hard to detect. With CRACI, the team created the first detailed maps of D in cellular tRNAs from mammals and plants, including those inside mitochondria.
Key discoveries:
Because D is common and conserved across species, and related enzymes are potentially linked to human disease, CRACI gives researchers a powerful platform to explore how these RNA marks shape cell function, stress responses, and health. The team plans to extend CRACI to primary cells and tissues to build a clear and comprehensive picture of D roles in disease contexts.
Journal reference
Ju, CW., Li, H., Jiang, B. et al. Quantitative CRACI reveals transcriptome-wide distribution of RNA dihydrouridine at base resolution. Nat Commun 16, 8863 (2025). https://doi.org/10.1038/s41467-025-63918-w
Read More
Prof. ZHAI Yuanliang, Associate Professor in the Division of Life Science (LIFS), has been awarded the Collaborative Research Project Grant (CRPG) under the Collaborative Research Fund (CRF) 2025/26. He was awarded HKD7.96 million for the project “Molecular Mechanisms of Replisome Coupling” in collaboration with The University of Hong Kong (HKU).
Read More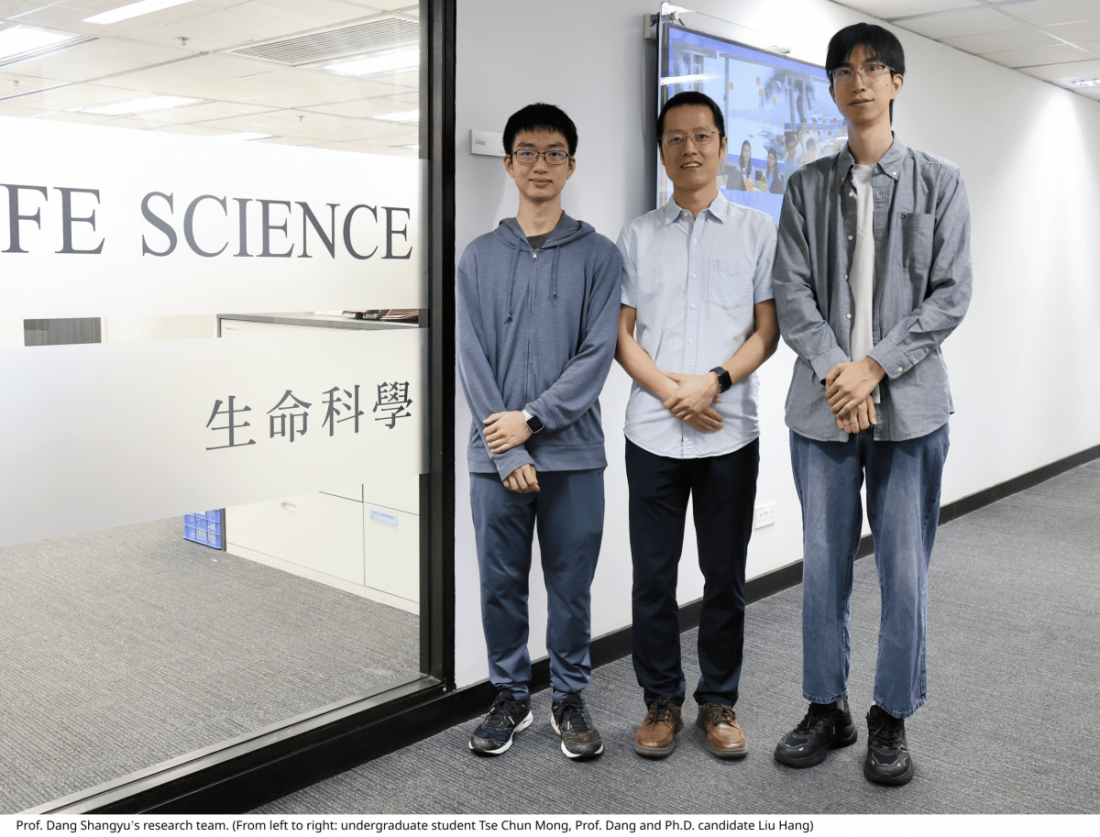
After four years of systematic investigation, Prof. Dang’s team bypassed the need for purification with detergents by directly generating vesicles containing the target protein from cell membranes. This approach produced samples suitable for cryo-EM imaging and structural studies. The team established a comprehensive workflow for the preparation, purification, and quality control of vesicle samples, making this method applicable to various membrane systems. To address the strong background signal and interference caused by the native membrane structure, they developed a micrograph-based sorting approach integrated with an artificial intelligence model to specifically isolate high-quality membrane protein particles. They successfully applied this method to multiple membrane protein systems, resolving the structure of the overexpressed AcrB protein in E. coli cell membranes at 3.9 Å resolution and the structure of the native respiratory chain complex III in porcine heart mitochondrial inner membranes at 3.0 Å resolution.
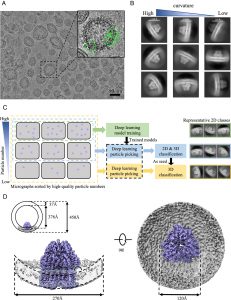
A scientific picture of Cryo-EM structure determination of AcrB in vesicle.
LIU Hang, a Ph.D. candidate from Prof. Dang’s team and the first author of the study, reflected: “Leveraging Prof. Dang’s multidisciplinary approach, our team has successfully developed a comprehensive system for in-situ structural studies of membrane proteins, encompassing both sample preparation and data processing, which overcomes challenges that were previously insurmountable. “
Compared to traditional detergent-based methods, this novel approach offers key advantages, including lower cost, simpler operation, and greater ease of use. Crucially, it preserves the native membrane environment and key lipid molecules, maintaining the protein’s natural conformation to the greatest extent possible. The method also demonstrates versatility, making it applicable to different membrane proteins across various species and cellular membrane structures. It promises to significantly reduce the workload for researchers, simplify the determination of membrane protein structures, and broaden the scope of cryo-EM structural biology.
“This vesicle-based platform preserves physiological lipid environments while eliminating the burdens of detergent screening,” explained Prof. Dang. “It provides an opportunity to study membrane proteins in their native environments. With further optimization, we aim to achieve structural proteomics of membrane proteins in specific biological membrane systems, such as mitochondria, under various physiological and pathological conditions, offering valuable insights into diseases.”
This research was published in the Proceedings of the National Academy of Sciences (PNAS). Prof. DANG Shangyu is the corresponding author, Ph.D. candidate LIU Hang is the first author, and undergraduate student TSE Chun Mong is the second author.
Repost from
Read More